UD researcher Kristi Kiick is part of a new study that provides insights into how disordered proteins form membrane-free liquid droplets, with implications for cell biology research, developing new disease treatments, and designing novel biomaterials.
Within all living cells are a diverse number of organelles—structures like ribosomes, mitochondria and the nucleus that have specialized functions. While the majority of these organelles are enclosed in a protective membrane, some are instead enclosed in isolated liquid droplets. The mechanisms by which these droplets form, a process called liquid-liquid phase separation, is a relatively new area of research that is hypothesized to drive many important biological processes such as transcription, X-chromosome inactivation, and tumorigenesis.
Now, a multi-institutional team of researchers have gained new insights into how disordered proteins form these membrane-free liquid droplets. Published in Nature Chemistry, these fundamental results could pave the way for a variety of future studies and applications, from insights on new disease treatments to unique bioengineered soft materials.

Kristi Kiick, a senior researcher in the College of Engineering’s Department of Biomedical Engineering and the Department of Materials Science and Engineering.
The study was led by Jeetain Mittal, Kenneth R. Hall professor in the Artie McFerrin Department of Chemical Engineering at Texas A&M University, and his research lab, in collaboration with Kristi Kiick, a senior researcher in the College of Engineering’s Department of Biomedical Engineering and the Department of Materials Science and Engineering, and assistant professor Ben Schuster from Rutgers University.
Liquid droplets in living cells were first observed in the germ cells of a common model organism known as Caenorhabditis elegans (C. elegans). In this study, a team of researchers found that, within the worm’s embryo, membrane-free structures called P granules serve essential reproductive functions. They also found that these P granules lacked membranes and could drip, join together or dissolve away. In short, they had some components that acted like liquids and others that were more solid, a property that solutions have after phase separation.
“There was a fundamental change in 2009 in thinking about cellular compartmentalization in terms of the emergence of droplet-like structures,” explained Mittal. “Most biologists began to accept that phase separation is not the exception but the rule with which biological cells compartmentalize functional units other than membrane-bound organelles.”
What wasn’t understood after that initial discovery was exactly how, out of the millions of proteins floating around in the cytoplasm, specific proteins could self-assemble into organized, functional droplets, and if and how that self-assembly capability was related to the droplet’s ability to phase separate.
“We still don’t have a very clear idea of which amino acids within the disordered regions provide the driving force for phase separation,” said Shiv Rekhi, a graduate student in Mittal’s laboratory and the paper’s first author. “We wanted to go outside of established rules, still show phase separation, and then quantify how each amino acid contributed to the process.”
To delve into this mechanism, the researchers used a synthetic disordered protein with amino acid sequences similar to those of a naturally occurring insect protein called resilin. Resilin, which has a unique ability to recover its original size and shape after stretching, has been studied by the Kiick group as a biomaterial because of its mechanical properties, as well as due to its ability to phase separate and isolate cells in certain regions of resilin-based biomaterials. The researchers aimed to predict the fundamental features of these proteins that can cause phase separation, then used this information to design next-generation biomaterials with finely tuned properties.
To determine the essential amino acid residues needed for phase separation to occur, the researchers created protein variants (by removing or adding a specific type of amino acid) to see if droplet condensation still occurred. Large-scale simulations were used by the Mittal group to evaluate how atomic interactions between individual amino acids translated to the formation of liquid droplets. The Kiick laboratories made the model proteins and verified phase separation results, with droplet characterization experiments conducted by the Schuster lab.
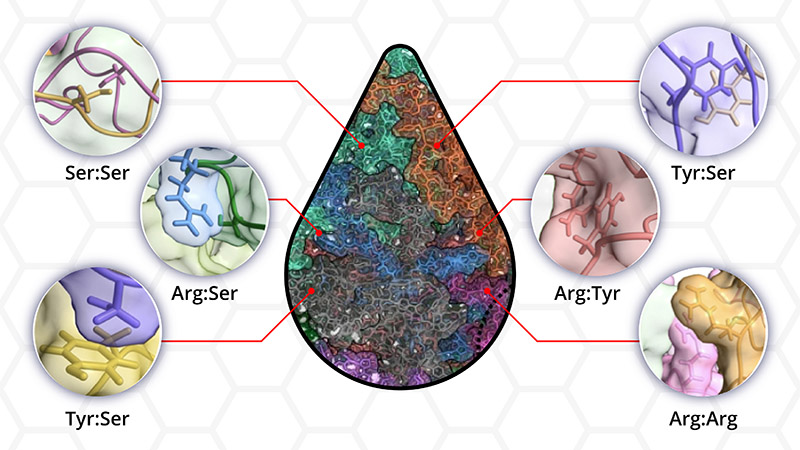
The results presented in this paper show that all but one of the 12 protein variants evaluated showed phase separation. These findings not only demonstrate that amino acids in droplets interact in many more ways than was currently recognized, it also underscores the presence of multiple interactions between the amino acid residues that make up the disordered protein.
“For a while, people in the field have assumed that a limited set of rules can describe droplet formation. We have shown that everything in the protein sequence matters,” said Mittal. “Our paper establishes that the molecular language of phase separation is much richer and more complex.”
“As materials scientists, we are excited about these results as they inform our design of new biomaterials,” said Kiick. “By controlling interactions between the disordered protein chains, we can control mechanical properties, microscale structures upon phase separation, and cell interactions in our materials. This will allow us in the future to design engineered scaffolds for cell-based therapies, drug delivery, and regenerative medicine applications.”
This research is funded by the National Science Foundation, the National Institute of General Medical Science of the National Institute of Health, and the Welch Foundation.
Adapted from a press release by Texas A&M Engineering.